✓ 24/7 automatic processing of online orders
✓ Knowledgeable and professional Product & Technical Support
✓ Fast and reliable (re)-ordering
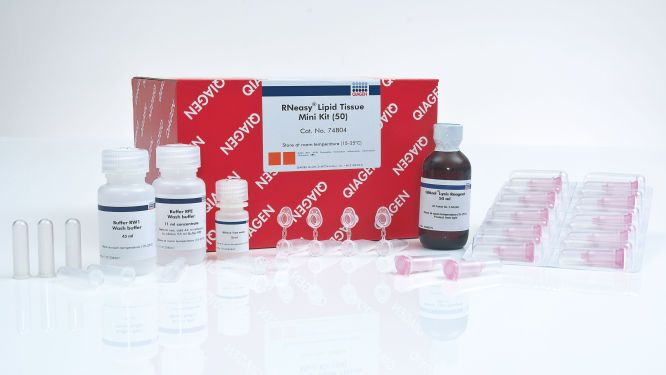
RNeasy Lipid Tissue Mini Kit (50)
Cat. No. / ID: 74804
✓ 24/7 automatic processing of online orders
✓ Knowledgeable and professional Product & Technical Support
✓ Fast and reliable (re)-ordering
Features
- Optimized lysis conditions for fatty tissues and other tissues
- High yields of total RNA without phenol contamination
- High-quality RNA for all downstream applications
Product Details
The RNeasy Lipid Tissue Mini Kit includes QIAzol Lysis Reagent for lysing fatty tissues and other types of tissue, and RNeasy spin columns for purifying up to 100 µg of high-quality RNA. The kit can be automated using the QIAcube Connect. Tissue samples can be conveniently stabilized using RNAprotect Tissue Reagent (nonfatty tissues only) or Allprotect Tissue Reagent, and efficiently disrupted using a TissueRuptor or TissueLyser system. For larger samples, the RNeasy Lipid Tissue Midi Kit (spin-column binding capacity of 1 mg RNA) is also available.
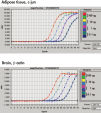
Performance
See figures
Principle
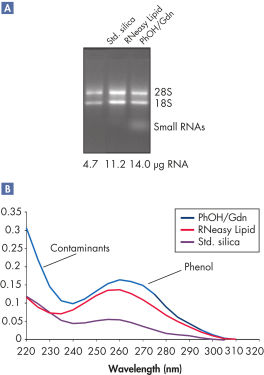
RNeasy Lipid Tissue Kits are optimized for use with fatty tissues, such as brain and adipose tissue. The convenient RNeasy Lipid Tissue protocol integrates optimized phenol/guanidine-based lysis with proven RNeasy purification for isolation of high yields of high-quality total RNA. The combination of organic extraction and chaotropic disruption contributes to the high lysis efficiency of QIAzol Lysis Reagent. Since the RNeasy procedures enrich for mRNA and other RNA species >200 nucleotides, the total RNA yield does not include 5S rRNA, tRNA, and other low-molecular-weight RNAs, which make up 15-20% of total cellular RNA.
Procedure
See figures
Applications
RNeasy Lipid Tissue Kits provide easy and efficient isolation of high-quality RNA for all downstream applications, including array analysis and real-time RT-PCR.
Supporting data and figures
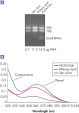
High yields of RNA without phenol carryover.
Specifications
| Features | Specifications |
|---|---|
| Applications | NGS, PCR, real-time PCR, microarray |
| Sample amount | 10–100 mg |
| Yield | 2.4–10 mg |
| Technology | Silica technology |
| Processing | Manual |
| Purification of total RNA, miRNA, poly A+ mRNA, DNA or protein | RNA |
| Elution volume | 30–100 µl |
| Main sample type | Fatty tissue samples |
| Time per run or per prep | 45 minutes |
| Format | Spin column |