Probe-based real-time을 one-step RT-PCR로 신속히 수행할 수 있습니다
Products
Features
- 60%까지 시간을 절약할 수 있는 보다 빠른 결과
- Sensitive detection of even low copy numbers
- 고감도 detection — low copy도 가능
- 광범위한 template에서의 정확하게 검출이 가능함
- 최적화된 master mix와 protocol
Product Details
Performance
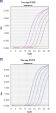
QuantiFast Probe RT-PCR Kits deliver highly sensitive results, outperforming other real-time RT-PCR kits (see figure " Sensitive One-step RT-PCR"). RT-PCR run times are reduced by up to 60% (see figure " Significantly reduced PCR times"), allowing you to achieve fast PCR results without compromising on RT-PCR performance (see figure " Faster results without compromising sensitivity"). You can also greatly increase your sample throughput or efficiently share a cycler with other users. With QuantiFast Probe RT-PCR Kits, you follow the same procedure whether you work with a standard or fast cycler.
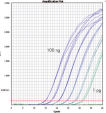
QuantiFast Probe RT-PCR Kits allow accurate quantification over a wide dynamic range (see figure " Comparable dynamic range"). Template dilutions of up to 6 logs can be reliably detected (see figure " Wide dynamic range and high sensitivity").
See figures
Principle
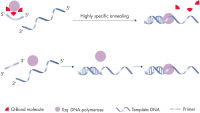
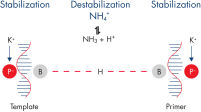
QuantiFast Probe RT-PCR Kits deliver highly sensitive and rapid results over a wide dynamic range on both standard and fast cyclers. The kits are designed for use with all types of sequence-specific probes, including hydrolysis probe detection (e.g., TaqMan and other dual-labeled probes) and FRET probes. The optimized QuantiFast RT Mix enables cDNA synthesis in just 10 minutes and a specially developed fast RT-PCR buffer contains the novel additive Q-Bond, which significantly reduces denaturation, annealing, and extension times (see figure " Fast primer annealing"). The RT-PCR buffer also contains a balanced combination of K+ and NH4+ ions, which promote specific primer annealing and enable high RT-PCR specificity and sensitivity (see figure " Specific primer annealing"). In addition, HotStarTaq Plus DNA Polymerase requires only 5 minutes at 95°C for activation and provides a stringent hot start, preventing the formation of nonspecific products.
| Component | Features and benefits | Benefits |
|---|---|---|
| HotStarTaq Plus DNA Polymerase | 5 min activation at 95ºC | Set up of qPCR reactions at room temperature |
| QuantiFast Probe RT-PCR Buffer | Balanced combination of NH4+ and K+ ions | Specific primer annealing ensures reliable PCR results |
| Unique Q-Bond additive | Faster PCR run times, enabling faster results and more reactions per day | |
| ROX dye† | Normalizes fluorescent signals on Applied Biosystems and, optionally, Agilent instruments | Precise quantification on cyclers that require ROX dye. Does not interfere with PCR on any real-time cycler |
| QuantiFast RT Mix | Separate solution added during PCR setup | Fast cDNA synthesis in just 10 minutes |
See figures
Procedure
QuantiFast Probe RT-PCR Kits contain ready-to-use master mixes that eliminate the need for optimization of reaction and cycling conditions. Simply add template RNA, primers, and probe to the master mix and follow the protocol in the handbook to get fast and reliable results on any real-time cycler. Kits are available with or without ROX passive reference dye in the master mix, enabling use on virtually any real-time cycler (see table). Due to the optimized ROX concentrations, detection of even low copy numbers is achieved through automatic data analysis.
| ROX dye | Kit | Compatible cyclers |
|---|---|---|
| Supplied in master mix | QuantiFast Probe RT-PCR Kit | All cyclers from Applied Biosystems except Applied Biosystems 7500 |
| Supplied in separate tube | QuantiFast Probe RT-PCR +ROX Vial Kit | Applied Biosystems 7500 and cyclers from Bio-Rad, Cepheid, Eppendorf, QIAGEN, Roche, Agilent, and other suppliers |
Applications
QuantiFast Probe RT-PCR Kits can be used for probe-based gene expression analysis of RNA targets on any real-time cycler. This includes instruments from Applied Biosystems, Bio-Rad, Cepheid, Eppendorf, Roche, and Agilent. For the Rotor-Gene Q and other Rotor-Gene cyclers, we recommend using the Rotor-Gene Probe RT-PCR Kit, which has been specially developed for fast cycling on these instruments.
Supporting data and figures
Significantly reduced RT-PCR times.

Specifications
| Features | Specifications |
|---|---|
| Applications | Probe-based, real-time RT-PCR |
| Single or multiplex | Single |
| Sample/target type | RNA |
| Real-time or endpoint | Real-time |
| Thermal cycler | All real-time cyclers (e.g. LC, RG, ABI) |
| Reaction type | Real-time one-step RT-PCR |
| SYBR Green I or sequence-specific probes | Sequence-specific probes |
| With or without ROX | Available with ROX in master mix and with ROX as a separate vial |